Protein Molecular Surface Calculator
The Protein Molecular Surface Calculator (ProMS) is a
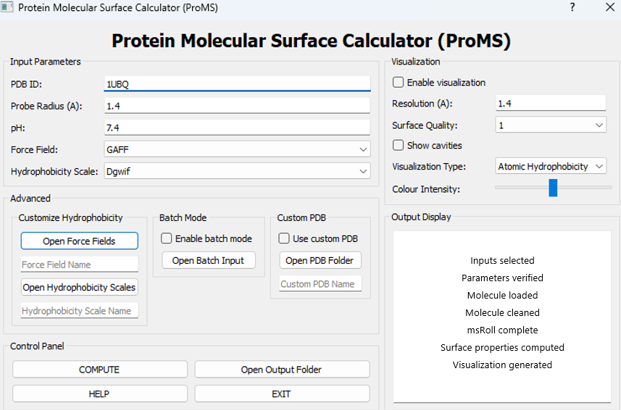
PyMOL-integrated computational tool designed to quantify and visualize physicochemical properties of molecular surfaces at atomic and residue resolution. ProMS enables users to analyze how proteins, nucleic acids, and small molecules interact with their environment by mapping key surface properties such as hydrophobicity and charge.
ProMS was initially released in 2022 and has since been updated with expanded functionality, improved stability, and enhanced integration within PyMOL. For access to the source code, please contact the lab.
Molecular Surfaces
Molecular surfaces play an integral role in many biological processes. They are involved in protein folding, protein-protein interactions, molecular recognition, molecular docking, signaling, and more. In 1983, Michael Connolly developed an algorithm to generate the geometrical molecular surface of any protein given as in input.

In simple terms, the “Connolly surface” is generated by rolling an imaginary sphere over the 3D structure of the molecule. The output surface is thus a function of the sphere’s radius (also referred to as the probe radius). To this day, Connolly’s algorithm remains the most popular analytical method for mapping protein surfaces.
In 1993, Connolly released software to encapsulate his algorithms for practical purposes. ProMS uses a portion of this software (notably msRoll.exe) as a step in calculating protein surface properties. Additionally, ProMS applies Nicolau’s work to investigate molecular surfaces at the atomic and amino acid resolution. This methodology provides a more accurate and higher-resolution description of the protein surface. Read more about it here.
ProMS Features
For a more detailed summary of ProMS’ features, please reference the user manual.
- Surface property calculations
- Computations derived from Connolly’s msRoll and Nicolau’s work on atomic hydrophobicities
- “Under the hood” – complexity and tediousness is hidden by the intuitive GUI
- Multiple output file formats, including a .pse (PyMOL session) file if visualization is enabled
- A sample of the output in tabular form can be found below
- Surface property calculations
- Choice of various input parameters
- PDB ID, probe radius (1.4-20A), pH, hydrophobicity scale
- Custom force fields and atom types
- Highly adaptive custom visualization module
- Choice of various input parameters
- Integration within PyMOL
- ProMS is an extension of Schrodinger’s PyMOL, an open-source molecular manipulation tool
- Easy-to-install plugin at the click of a few buttons
- Integration within PyMOL
- Batch mode
- Enables the analysis of several proteins / different parameters at once
- Batch mode
- Custom PDB integration
- Supports PDBs that have been modified by the user or that have not yet been published to an online database
- Custom PDB integration
ProMS Sample Output
Below is a sample of the properties calculated by ProMS, aggregated into a table. The analyzed proteins correspond to the same proteins chosen for the BAD project. These properties were calculated using Nicolau’s Method 1 and the “Dgwif” hydrophobicity scale. There are four copies of each PDB because the probe radius (1.4A vs 20A) and the resolution (atomic vs amino acid) were changed (see last two columns).
The majority of the output columns are self-explanatory. However, some of the secondary properties, such as density, specific density, and area extent are less intuitive.
-
- Density = total charge or hydrophobicity / total area
- Specific density = total charge or hydrophobicity / total charged area or hydrophobic area
- Area extent = charged or hydrophobic area / total area
- All areas are in units of square Angstroms
| ID | PDB | Probe Radius | Resolution | Total Surface Area | Positive Area | Negative Area | Hydrophobic Area | Hydrophilic Area | Total Positive Charge | Total Negative Charge | Total Hydrophobicity | Total Hydrophilicity | Positive Charge Density | Positive Charge Specific Density | Negative Charge Density | Negative Charge Specific Density | Positive Area Extent | Negative Area Extent | Hydrophobic Density | Hydrophobic Specific Density | Hydrophilic Density | Hydrophilic Specific Density | Hydrophobic Area Extent | Hydrophilic Area Extent |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 1A4V | 1.4 | AT | 7834 | 827 | 4150 | 2050 | 2927 | 34 | -87 | -252 | 114 | 0.0044 | 0.0412 | -0.0112 | -0.0211 | 0.1056 | 0.5297 | -0.0321 | -0.1228 | 0.0146 | 0.0390 | 0.2617 | 0.3736 |
| 2 | 1A4V | 20.0 | AT | 4690 | 1126 | 3416 | 2878 | 1664 | 42 | -65 | -239 | 21 | 0.0089 | 0.0373 | -0.0139 | -0.0191 | 0.2400 | 0.7284 | -0.0509 | -0.0830 | 0.0045 | 0.0127 | 0.6136 | 0.3548 |
| 3 | 1AO6 | 1.4 | AA | 55064 | 9597 | 45467 | 45374 | 8150 | 28 | -120 | -387 | 49 | 0.001 | 0.003 | -0.002 | -0.003 | 0.174 | 0.826 | -0.007 | -0.009 | 0.001 | 0.006 | 0.824 | 0.148 |
| 4 | 1AO6 | 1.4 | AT | 55064 | 9597 | 45467 | 22081 | 32983 | 395 | -974 | -2980 | 1465 | 0.007 | 0.041 | -0.018 | -0.021 | 0.174 | 0.826 | -0.054 | -0.135 | 0.027 | 0.044 | 0.401 | 0.599 |
| 5 | 1AO6 | 20.0 | AA | 26983 | 6381 | 20601 | 26756 | 179 | 43 | -94 | -301 | 1 | 0.002 | 0.007 | -0.003 | -0.005 | 0.236 | 0.764 | -0.011 | -0.011 | 0 | 0.006 | 0.992 | 0.007 |
| 6 | 1AO6 | 20.0 | AT | 26983 | 6094 | 20451 | 18764 | 7781 | 280 | -413 | -1677 | 96 | 0.01 | 0.046 | -0.015 | -0.02 | 0.226 | 0.758 | -0.062 | -0.089 | 0.004 | 0.012 | 0.695 | 0.288 |
| 7 | 1B0L | 1.4 | AA | 28019 | 6418 | 21601 | 23691 | 3892 | 13 | -59 | -162 | 24 | 0 | 0.002 | -0.002 | -0.003 | 0.229 | 0.771 | -0.006 | -0.007 | 0.001 | 0.006 | 0.846 | 0.139 |
| 8 | 1B0L | 1.4 | AT | 28019 | 6418 | 21601 | 12789 | 15230 | 181 | -458 | -1672 | 789 | 0.006 | 0.028 | -0.016 | -0.021 | 0.229 | 0.771 | -0.06 | -0.131 | 0.028 | 0.052 | 0.456 | 0.544 |
| 9 | 1B0L | 20.0 | AA | 16486 | 5756 | 10729 | 16302 | 130 | 17 | -42 | -140 | 0 | 0.001 | 0.003 | -0.003 | -0.004 | 0.349 | 0.651 | -0.009 | -0.009 | 0 | 0.007 | 0.989 | 0.008 |
| 10 | 1B0L | 20.0 | AT | 16486 | 5632 | 10260 | 10856 | 5037 | 118 | -183 | -897 | 66 | 0.007 | 0.021 | -0.011 | -0.018 | 0.342 | 0.622 | -0.054 | -0.083 | 0.004 | 0.013 | 0.658 | 0.306 |
| 11 | 1BJ5 | 1.4 | AT | 28109 | 4632 | 23148 | 10537 | 17244 | 183 | -477 | -1283 | 674 | 0.0065 | 0.0395 | -0.0170 | -0.0206 | 0.1648 | 0.8235 | -0.0456 | -0.1218 | 0.0240 | 0.0391 | 0.3748 | 0.6135 |
| 12 | 1BJ5 | 20.0 | AT | 15875 | 2496 | 12338 | 9348 | 5486 | 90 | -239 | -759 | 50 | 0.0057 | 0.0362 | -0.0151 | -0.0194 | 0.1572 | 0.7772 | -0.0478 | -0.0812 | 0.0031 | 0.0091 | 0.5888 | 0.3456 |
| 13 | 1BUW | 1.4 | AA | 25381 | 4068 | 21272 | 20142 | 5065 | 11 | -52 | -107 | 32 | 0 | 0.003 | -0.002 | -0.002 | 0.16 | 0.838 | -0.004 | -0.005 | 0.001 | 0.006 | 0.794 | 0.2 |
| 14 | 1BUW | 1.4 | AT | 25381 | 4068 | 21272 | 8939 | 16401 | 156 | -407 | -1198 | 835 | 0.006 | 0.038 | -0.016 | -0.019 | 0.16 | 0.838 | -0.047 | -0.134 | 0.033 | 0.051 | 0.352 | 0.646 |
| 15 | 1BUW | 20.0 | AA | 12702 | 3295 | 9407 | 11546 | 1156 | 18 | -35 | -72 | 6 | 0.001 | 0.005 | -0.003 | -0.004 | 0.259 | 0.741 | -0.006 | -0.006 | 0.001 | 0.006 | 0.909 | 0.091 |
| 16 | 1BUW | 20.0 | AT | 12702 | 3283 | 9374 | 6547 | 6110 | 129 | -167 | -601 | 77 | 0.01 | 0.039 | -0.013 | -0.018 | 0.258 | 0.738 | -0.047 | -0.092 | 0.006 | 0.013 | 0.515 | 0.481 |
| 17 | 1CF3 | 1.4 | AA | 20897 | 4096 | 16801 | 16586 | 4244 | 7 | -47 | -109 | 25 | 0 | 0.002 | -0.002 | -0.003 | 0.196 | 0.804 | -0.005 | -0.007 | 0.001 | 0.006 | 0.794 | 0.203 |
| 18 | 1CF3 | 1.4 | AT | 20897 | 4096 | 16801 | 9462 | 11435 | 121 | -358 | -1237 | 595 | 0.006 | 0.03 | -0.017 | -0.021 | 0.196 | 0.804 | -0.059 | -0.131 | 0.029 | 0.052 | 0.453 | 0.547 |
| 19 | 1CF3 | 20.0 | AA | 12549 | 3100 | 9382 | 11900 | 648 | 11 | -42 | -104 | 4 | 0.001 | 0.004 | -0.003 | -0.005 | 0.247 | 0.748 | -0.008 | -0.009 | 0 | 0.006 | 0.948 | 0.052 |
| 20 | 1CF3 | 20.0 | AT | 12549 | 2994 | 9221 | 9049 | 3166 | 85 | -173 | -764 | 40 | 0.007 | 0.028 | -0.014 | -0.019 | 0.239 | 0.735 | -0.061 | -0.084 | 0.003 | 0.013 | 0.721 | 0.252 |
| 21 | 1CHO | 1.4 | AT | 11913 | 1636 | 5805 | 3375 | 4066 | 49 | -125 | -367 | 192 | 0.0041 | 0.0301 | -0.0105 | -0.0216 | 0.1373 | 0.4873 | -0.0308 | -0.1089 | 0.0161 | 0.0472 | 0.2833 | 0.3413 |
| 22 | 1CHO | 20.0 | AT | 7746 | 1705 | 5630 | 4736 | 2599 | 46 | -91 | -310 | 18 | 0.0060 | 0.0271 | -0.0118 | -0.0162 | 0.2201 | 0.7269 | -0.0400 | -0.0654 | 0.0023 | 0.0069 | 0.6114 | 0.3356 |
| 23 | 1GPE | 1.4 | AT | 31439 | 4366 | 17130 | 9387 | 12109 | 157 | -366 | -1105 | 484 | 0.0050 | 0.0360 | -0.0116 | -0.0214 | 0.1389 | 0.5449 | -0.0351 | -0.1177 | 0.0154 | 0.0400 | 0.2986 | 0.3852 |
| 24 | 1GPE | 20.0 | AT | 20175 | 5125 | 14374 | 11437 | 8061 | 139 | -241 | -810 | 59 | 0.0069 | 0.0272 | -0.0119 | -0.0167 | 0.2540 | 0.7124 | -0.0402 | -0.0708 | 0.0029 | 0.0074 | 0.5669 | 0.3996 |
| 25 | 1HLS | 1.4 | AT | 3137 | 590 | 2548 | 1859 | 1278 | 15 | -51 | 76 | -152 | 0.0047 | 0.0248 | -0.0162 | -0.0199 | 0.1880 | 0.8120 | 0.0243 | 0.0410 | -0.0483 | -0.1187 | 0.5927 | 0.4073 |
| 26 | 1HLS | 20.0 | AT | 2648 | 472 | 2162 | 1615 | 1018 | 6 | -37 | -106 | 12 | 0.0021 | 0.0121 | -0.0139 | -0.0170 | 0.1781 | 0.8163 | -0.0402 | -0.0659 | 0.0045 | 0.0117 | 0.6100 | 0.3845 |
| 27 | 1HML | 1.4 | AA | 6097 | 1103 | 4993 | 4271 | 1694 | 2 | -13 | -36 | 8 | 0 | 0.003 | -0.002 | -0.003 | 0.181 | 0.819 | -0.006 | -0.009 | 0.001 | 0.005 | 0.701 | 0.278 |
| 28 | 1HML | 1.4 | AT | 6097 | 1103 | 4993 | 2581 | 3515 | 43 | -107 | -354 | 189 | 0.007 | 0.04 | -0.018 | -0.022 | 0.181 | 0.819 | -0.058 | -0.137 | 0.031 | 0.054 | 0.423 | 0.577 |
| 29 | 1HML | 20.0 | AA | 4692 | 1112 | 3536 | 3805 | 886 | 5 | -13 | -38 | 4 | 0.001 | 0.005 | -0.003 | -0.004 | 0.237 | 0.754 | -0.008 | -0.01 | 0.001 | 0.005 | 0.811 | 0.189 |
| 30 | 1HML | 20.0 | AT | 4692 | 1100 | 3371 | 2940 | 1531 | 44 | -61 | -267 | 20 | 0.009 | 0.04 | -0.013 | -0.018 | 0.235 | 0.718 | -0.057 | -0.091 | 0.004 | 0.013 | 0.627 | 0.326 |
| 31 | 1HRC | 1.4 | AA | 5479 | 1038 | 4440 | 4573 | 830 | 3 | -11 | -34 | 4 | 0.001 | 0.004 | -0.002 | -0.003 | 0.19 | 0.81 | -0.006 | -0.008 | 0.001 | 0.006 | 0.835 | 0.152 |
| 32 | 1HRC | 1.4 | AT | 5479 | 1038 | 4440 | 2311 | 3167 | 42 | -92 | -273 | 104 | 0.008 | 0.041 | -0.017 | -0.021 | 0.19 | 0.81 | -0.05 | -0.118 | 0.019 | 0.033 | 0.422 | 0.578 |
| 33 | 1HRC | 20.0 | AA | 3973 | 1288 | 2639 | 3831 | 141 | 8 | -10 | -33 | 0 | 0.002 | 0.007 | -0.003 | -0.004 | 0.324 | 0.664 | -0.008 | -0.009 | 0 | 0.003 | 0.964 | 0.036 |
| 34 | 1HRC | 20.0 | AT | 3973 | 1279 | 2605 | 2477 | 1406 | 56 | -46 | -191 | 11 | 0.014 | 0.044 | -0.012 | -0.018 | 0.322 | 0.656 | -0.048 | -0.077 | 0.003 | 0.008 | 0.624 | 0.354 |
| 35 | 1IGT | 1.4 | AA | 59573 | 12441 | 47131 | 47611 | 10947 | 24 | -132 | -279 | 61 | 0 | 0.002 | -0.002 | -0.003 | 0.209 | 0.791 | -0.005 | -0.006 | 0.001 | 0.006 | 0.799 | 0.184 |
| 36 | 1IGT | 1.4 | AT | 59573 | 12441 | 47131 | 26382 | 33191 | 343 | -984 | -3121 | 1799 | 0.006 | 0.028 | -0.017 | -0.021 | 0.209 | 0.791 | -0.052 | -0.118 | 0.03 | 0.054 | 0.443 | 0.557 |
| 37 | 1IGT | 20.0 | AA | 37314 | 9291 | 27977 | 32794 | 3493 | 35 | -111 | -215 | 20 | 0.001 | 0.004 | -0.003 | -0.004 | 0.249 | 0.75 | -0.006 | -0.007 | 0.001 | 0.006 | 0.879 | 0.094 |
| 38 | 1IGT | 20.0 | AT | 37314 | 8820 | 27129 | 23247 | 12702 | 229 | -479 | -1502 | 160 | 0.006 | 0.026 | -0.013 | -0.018 | 0.236 | 0.727 | -0.04 | -0.065 | 0.004 | 0.013 | 0.623 | 0.34 |
| 39 | 1M1J | 1.4 | AT | 88145 | 18119 | 68552 | 37073 | 49597 | 580 | -1441 | -4183 | 1947 | 0.0066 | 0.0320 | -0.0164 | -0.0210 | 0.2056 | 0.7777 | -0.0475 | -0.1128 | 0.0221 | 0.0393 | 0.4206 | 0.5627 |
| 40 | 1M1J | 20.0 | AT | 55493 | 15143 | 38709 | 33059 | 20793 | 400 | -673 | -2240 | 222 | 0.0072 | 0.0264 | -0.0121 | -0.0174 | 0.2729 | 0.6976 | -0.0404 | -0.0677 | 0.0040 | 0.0107 | 0.5957 | 0.3747 |
| 41 | 1PIF | 1.4 | AA | 17193 | 4104 | 13089 | 13474 | 3528 | 7 | -36 | -79 | 27 | 0 | 0.002 | -0.002 | -0.003 | 0.239 | 0.761 | -0.005 | -0.006 | 0.002 | 0.008 | 0.784 | 0.205 |
| 42 | 1PIF | 1.4 | AT | 17193 | 4104 | 13089 | 7955 | 9237 | 99 | -279 | -919 | 388 | 0.006 | 0.024 | -0.016 | -0.021 | 0.239 | 0.761 | -0.053 | -0.116 | 0.023 | 0.042 | 0.463 | 0.537 |
| 43 | 1PIF | 20.0 | AA | 11355 | 3023 | 8331 | 10058 | 1242 | 7 | -31 | -63 | 11 | 0.001 | 0.003 | -0.003 | -0.004 | 0.266 | 0.734 | -0.006 | -0.006 | 0.001 | 0.01 | 0.886 | 0.109 |
| 44 | 1PIF | 20.0 | AT | 11355 | 2954 | 8058 | 6666 | 4346 | 65 | -133 | -469 | 46 | 0.006 | 0.022 | -0.012 | -0.017 | 0.26 | 0.71 | -0.041 | -0.07 | 0.004 | 0.011 | 0.587 | 0.383 |
| 45 | 2CHA | 1.4 | AA | 19010 | 4537 | 14473 | 15317 | 3341 | 8 | -41 | -74 | 22 | 0 | 0.002 | -0.002 | -0.003 | 0.239 | 0.761 | -0.004 | -0.005 | 0.001 | 0.007 | 0.806 | 0.176 |
| 46 | 2CHA | 1.4 | AT | 19010 | 4537 | 14473 | 8549 | 10461 | 118 | -299 | -935 | 519 | 0.006 | 0.026 | -0.016 | -0.021 | 0.239 | 0.761 | -0.049 | -0.109 | 0.027 | 0.05 | 0.45 | 0.55 |
| 47 | 2CHA | 20.0 | AA | 11137 | 4534 | 6490 | 10388 | 468 | 18 | -27 | -63 | 2 | 0.002 | 0.004 | -0.003 | -0.004 | 0.407 | 0.583 | -0.006 | -0.006 | 0 | 0.006 | 0.933 | 0.042 |
| 48 | 2CHA | 20.0 | AT | 11137 | 4276 | 6475 | 7445 | 3305 | 116 | -106 | -472 | 15 | 0.01 | 0.027 | -0.01 | -0.016 | 0.384 | 0.581 | -0.042 | -0.064 | 0.001 | 0.005 | 0.669 | 0.297 |
| 49 | 2LYZ | 1.4 | AA | 5796 | 1616 | 4179 | 4667 | 1001 | 2 | -11 | -25 | 7 | 0 | 0.002 | -0.002 | -0.003 | 0.279 | 0.721 | -0.004 | -0.005 | 0.001 | 0.007 | 0.805 | 0.173 |
| 50 | 2LYZ | 1.4 | AT | 5796 | 1616 | 4179 | 2658 | 3138 | 36 | -84 | -335 | 196 | 0.006 | 0.023 | -0.015 | -0.02 | 0.279 | 0.721 | -0.058 | -0.126 | 0.034 | 0.063 | 0.459 | 0.541 |
| 51 | 2LYZ | 20.0 | AA | 4489 | 2014 | 2449 | 4192 | 251 | 4 | -9 | -23 | 1 | 0.001 | 0.002 | -0.002 | -0.004 | 0.449 | 0.546 | -0.005 | -0.006 | 0 | 0.006 | 0.934 | 0.056 |
| 52 | 2LYZ | 20.0 | AT | 4489 | 2014 | 2372 | 3164 | 1223 | 29 | -35 | -243 | 13 | 0.006 | 0.014 | -0.008 | -0.015 | 0.449 | 0.528 | -0.054 | -0.077 | 0.003 | 0.011 | 0.705 | 0.272 |
| 53 | 2RCJ | 1.4 | AT | 312775 | 0 | 312775 | 0 | 312775 | 0 | -4805 | 0 | 3227 | 0.0000 | 0.0000 | -0.0154 | -0.0154 | 0.0000 | 1.0000 | 0.0000 | 0.0000 | 0.0103 | 0.0103 | 0.0000 | 1.0000 |
| 54 | 2RCJ | 20.0 | AT | 168174 | 0 | 36468 | 0 | 36468 | 0 | -376 | 0 | 235 | 0.0000 | 0.0000 | -0.0022 | -0.0103 | 0.0000 | 0.2168 | 0.0000 | 0.0000 | 0.0014 | 0.0064 | 0.0000 | 0.2168 |
| 55 | 3BLG | 1.4 | AA | 7470 | 1277 | 6192 | 5829 | 1589 | 3 | -16 | -48 | 7 | 0.001 | 0.003 | -0.002 | -0.003 | 0.171 | 0.829 | -0.006 | -0.008 | 0.001 | 0.005 | 0.78 | 0.213 |
| 56 | 3BLG | 1.4 | AT | 7470 | 1277 | 6192 | 3118 | 4351 | 53 | -133 | -430 | 224 | 0.007 | 0.042 | -0.018 | -0.022 | 0.171 | 0.829 | -0.058 | -0.138 | 0.03 | 0.051 | 0.417 | 0.583 |
| 57 | 3BLG | 20.0 | AA | 5417 | 1284 | 4068 | 4961 | 456 | 7 | -16 | -50 | 2 | 0.001 | 0.006 | -0.003 | -0.004 | 0.237 | 0.751 | -0.009 | -0.01 | 0 | 0.004 | 0.916 | 0.084 |
| 58 | 3BLG | 20.0 | AT | 5417 | 1281 | 3989 | 3385 | 1885 | 49 | -75 | -306 | 22 | 0.009 | 0.039 | -0.014 | -0.019 | 0.236 | 0.736 | -0.056 | -0.09 | 0.004 | 0.012 | 0.625 | 0.348 |
| 59 | 3GHG | 1.4 | AA | 91878 | 19446 | 72356 | 69525 | 20575 | 41 | -185 | -457 | 117 | 0 | 0.002 | -0.002 | -0.003 | 0.212 | 0.788 | -0.005 | -0.007 | 0.001 | 0.006 | 0.757 | 0.224 |
| 60 | 3GHG | 1.4 | AT | 91878 | 19446 | 72356 | 37185 | 54617 | 599 | -1487 | -4754 | 2689 | 0.007 | 0.031 | -0.016 | -0.021 | 0.212 | 0.788 | -0.052 | -0.128 | 0.029 | 0.049 | 0.405 | 0.594 |
| 61 | 3GHG | 20.0 | AA | 55851 | 18845 | 36805 | 44127 | 10904 | 66 | -130 | -332 | 65 | 0.001 | 0.004 | -0.002 | -0.004 | 0.337 | 0.659 | -0.006 | -0.008 | 0.001 | 0.006 | 0.79 | 0.195 |
| 62 | 3GHG | 20.0 | AT | 55851 | 18001 | 35344 | 33742 | 19603 | 444 | -588 | -2640 | 248 | 0.008 | 0.025 | -0.011 | -0.017 | 0.322 | 0.633 | -0.047 | -0.078 | 0.004 | 0.013 | 0.604 | 0.351 |
| 63 | 3M7P | 1.4 | AA | 15314 | 3187 | 12126 | 11479 | 3425 | 4 | -31 | -69 | 20 | 0 | 0.002 | -0.002 | -0.003 | 0.208 | 0.792 | -0.005 | -0.006 | 0.001 | 0.006 | 0.75 | 0.224 |
| 64 | 3M7P | 1.4 | AT | 15314 | 3187 | 12126 | 6743 | 8571 | 86 | -248 | -874 | 551 | 0.006 | 0.027 | -0.016 | -0.02 | 0.208 | 0.792 | -0.057 | -0.13 | 0.036 | 0.064 | 0.44 | 0.56 |
| 65 | 3M7P | 20.0 | AA | 10102 | 2600 | 7501 | 7723 | 2270 | 4 | -26 | -54 | 13 | 0 | 0.002 | -0.003 | -0.003 | 0.257 | 0.743 | -0.005 | -0.007 | 0.001 | 0.006 | 0.765 | 0.225 |
| 66 | 3M7P | 20.0 | AT | 10102 | 2212 | 7096 | 5691 | 3618 | 30 | -115 | -466 | 45 | 0.003 | 0.014 | -0.011 | -0.016 | 0.219 | 0.702 | -0.046 | -0.082 | 0.004 | 0.013 | 0.563 | 0.358 |
| 67 | 3RGK | 1.4 | AA | 7215 | 1326 | 5888 | 6006 | 1208 | 4 | -15 | -48 | 6 | 0.001 | 0.003 | -0.002 | -0.003 | 0.184 | 0.816 | -0.007 | -0.008 | 0.001 | 0.006 | 0.832 | 0.168 |
| 68 | 3RGK | 1.4 | AT | 7215 | 1326 | 5888 | 2869 | 4346 | 57 | -122 | -353 | 170 | 0.008 | 0.043 | -0.017 | -0.021 | 0.184 | 0.816 | -0.049 | -0.123 | 0.024 | 0.039 | 0.398 | 0.602 |
| 69 | 3RGK | 20.0 | AA | 4933 | 1814 | 3068 | 4841 | 91 | 10 | -11 | -47 | 0 | 0.002 | 0.006 | -0.002 | -0.004 | 0.368 | 0.622 | -0.01 | -0.01 | 0 | 0.006 | 0.981 | 0.019 |
| 70 | 3RGK | 20.0 | AT | 4933 | 1732 | 2911 | 3131 | 1512 | 67 | -56 | -311 | 19 | 0.014 | 0.039 | -0.011 | -0.019 | 0.351 | 0.59 | -0.063 | -0.099 | 0.004 | 0.013 | 0.635 | 0.307 |
| 71 | 4ACQ | 1.4 | AA | 240174 | 45355 | 194818 | 182574 | 53749 | 82 | -489 | -1171 | 322 | 0 | 0.002 | -0.002 | -0.003 | 0.189 | 0.811 | -0.005 | -0.006 | 0.001 | 0.006 | 0.76 | 0.224 |
| 72 | 4ACQ | 1.4 | AT | 240174 | 45355 | 194818 | 93697 | 146476 | 1324 | -3835 | -12210 | 9354 | 0.006 | 0.029 | -0.016 | -0.02 | 0.189 | 0.811 | -0.051 | -0.13 | 0.039 | 0.064 | 0.39 | 0.61 |
| 73 | 4ACQ | 20.0 | AA | 100260 | 22346 | 77913 | 82990 | 16695 | 86 | -252 | -685 | 116 | 0.001 | 0.004 | -0.003 | -0.003 | 0.223 | 0.777 | -0.007 | -0.008 | 0.001 | 0.007 | 0.828 | 0.167 |
| 74 | 4ACQ | 20.0 | AT | 100260 | 21405 | 71478 | 51765 | 41118 | 525 | -1106 | -3973 | 465 | 0.005 | 0.025 | -0.011 | -0.015 | 0.214 | 0.713 | -0.04 | -0.077 | 0.005 | 0.011 | 0.516 | 0.41 |
| 75 | 4ARX | 1.4 | AT | 85337 | 19583 | 65754 | 45815 | 39522 | 341 | -1317 | 1506 | -4382 | 0.0040 | 0.0174 | -0.0154 | -0.0200 | 0.2295 | 0.7705 | 0.0176 | 0.0329 | -0.0513 | -0.1109 | 0.5369 | 0.4631 |
| 76 | 4ARX | 20.0 | AT | 27747 | 7228 | 19255 | 16552 | 9931 | 140 | -327 | -1252 | 134 | 0.0050 | 0.0190 | -0.0120 | -0.0170 | 0.2610 | 0.6940 | -0.0450 | -0.0760 | 0.0050 | 0.0130 | 0.5970 | 0.3580 |
| 77 | 4INS | 1.4 | AA | 5265 | 954 | 4311 | 3461 | 1465 | 1 | -10 | -21 | 7 | 0 | 0.002 | -0.002 | -0.002 | 0.181 | 0.819 | -0.004 | -0.006 | 0.001 | 0.005 | 0.657 | 0.278 |
| 78 | 4INS | 1.4 | AT | 5265 | 954 | 4311 | 2191 | 3074 | 25 | -84 | -251 | 118 | 0.005 | 0.027 | -0.016 | -0.02 | 0.181 | 0.819 | -0.048 | -0.115 | 0.022 | 0.038 | 0.416 | 0.584 |
| 79 | 4INS | 20.0 | AA | 3944 | 437 | 2888 | 2750 | 1028 | 0 | -7 | -18 | 5 | 0 | 0.002 | -0.002 | -0.003 | 0.111 | 0.732 | -0.005 | -0.007 | 0.001 | 0.005 | 0.697 | 0.261 |
| 80 | 4INS | 20.0 | AT | 3944 | 379 | 2842 | 1300 | 1921 | 8 | -38 | -91 | 29 | 0.002 | 0.023 | -0.01 | -0.013 | 0.096 | 0.721 | -0.023 | -0.07 | 0.008 | 0.016 | 0.33 | 0.487 |
| 81 | 4W8J | 1.4 | AA | 43779 | 9901 | 33877 | 34526 | 9252 | 15 | -90 | -232 | 58 | 0 | 0.002 | -0.002 | -0.003 | 0.226 | 0.774 | -0.005 | -0.007 | 0.001 | 0.006 | 0.789 | 0.211 |
| 82 | 4W8J | 1.4 | AT | 43779 | 9901 | 33877 | 19615 | 24164 | 248 | -711 | -2497 | 1305 | 0.006 | 0.025 | -0.016 | -0.021 | 0.226 | 0.774 | -0.057 | -0.127 | 0.03 | 0.054 | 0.448 | 0.552 |
| 83 | 4W8J | 20.0 | AA | 27746 | 7862 | 19884 | 24167 | 3579 | 19 | -71 | -197 | 27 | 0.001 | 0.003 | -0.003 | -0.004 | 0.283 | 0.717 | -0.007 | -0.008 | 0.001 | 0.008 | 0.871 | 0.129 |
| 84 | 4W8J | 20.0 | AT | 27746 | 7228 | 19254 | 16551 | 9931 | 139 | -326 | -1252 | 133 | 0.005 | 0.019 | -0.012 | -0.017 | 0.261 | 0.694 | -0.045 | -0.076 | 0.005 | 0.013 | 0.597 | 0.358 |
| 85 | 6KXS | 1.4 | AA | 136902 | 28291 | 108610 | 110451 | 24976 | 47 | -292 | -676 | 138 | 0 | 0.002 | -0.002 | -0.003 | 0.207 | 0.793 | -0.005 | -0.006 | 0.001 | 0.006 | 0.807 | 0.182 |
| 86 | 6KXS | 1.4 | AT | 136902 | 28291 | 108610 | 58733 | 78168 | 744 | -2207 | -7235 | 4458 | 0.005 | 0.026 | -0.016 | -0.02 | 0.207 | 0.793 | -0.053 | -0.123 | 0.033 | 0.057 | 0.429 | 0.571 |
| 87 | 6KXS | 20.0 | AA | 66836 | 14854 | 51982 | 60890 | 5297 | 33 | -190 | -457 | 29 | 0.001 | 0.002 | -0.003 | -0.004 | 0.222 | 0.778 | -0.007 | -0.008 | 0 | 0.006 | 0.911 | 0.079 |
| 88 | 6KXS | 20.0 | AT | 66836 | 13663 | 50559 | 37792 | 26429 | 204 | -820 | -2731 | 320 | 0.003 | 0.015 | -0.012 | -0.016 | 0.204 | 0.756 | -0.041 | -0.072 | 0.005 | 0.012 | 0.565 | 0.395 |
| 89 | 6TAV | 1.4 | AT | 240174 | 45355 | 194818 | 93697 | 146476 | 1324 | -3835 | -12210 | 9354 | 0.0055 | 0.0292 | -0.0160 | -0.0197 | 0.1888 | 0.8112 | -0.0508 | -0.1303 | 0.0389 | 0.0639 | 0.3901 | 0.6099 |
| 90 | 6TAV | 20.0 | AT | 100261 | 21406 | 71478 | 51766 | 12702 | 525 | -1106 | -3974 | 466 | 0.0052 | 0.0245 | -0.0110 | -0.0155 | 0.2135 | 0.7129 | -0.0396 | -0.0768 | 0.0046 | 0.0367 | 0.5163 | 0.1267 |
Additional Output
In addition to the properties above, ProMS also outputs detailed area calculations and the solvent-excluded volume via Connolly’s msRoll. Furthermore, using the protein’s molecular weight from literature, an atomic volume can also be calculated. Lastly, sphericity, monolayer volume density, and monolayer weight can all be computed from the aforementioned values. The results can be found in the table below.
| ID | Protein | PDB | Probe Radius | Molecular Weight | Atomic Volume | Contact Area | Reentrant Area | Molecular Area | Accessible Area | Solvent-Excluded Volume | Sphericity | Monolayer Volume Density | Monolayer Weight |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | Alpha-2-macroglobulin | 6TAV | 1.4 | 666.20 | 746144 | 70808 | 169367 | 240174 | 225522 | 669383 | 0.1541 | 0.0922 | 1.3815 |
| 2 | Alpha-2-macroglobulin | 6TAV | 20.0 | 666.20 | 746144 | 1262 | 89101 | 90363 | 141849 | 1405237 | 0.6714 | 0.5628 | 5.1454 |
| 3 | Alpha-amylase | 1PIF | 1.4 | 55.47 | 62126 | 5451 | 11742 | 17193 | 17508 | 64675 | 0.4532 | 0.4194 | 2.4863 |
| 4 | Alpha-amylase | 1PIF | 20.0 | 55.47 | 62126 | 218 | 11138 | 11356 | 33308 | 90497 | 0.8584 | 0.6000 | 2.8432 |
| 5 | Alpha-chymotrypsin | 1CHO | 1.4 | 31.29 | 35045 | 3795 | 7602 | 11396 | 12193 | 35672 | 0.4598 | 0.4256 | 2.1162 |
| 6 | Alpha-chymotrypsin | 1CHO | 20.0 | 31.29 | 35045 | 170 | 7576 | 7746 | 26385 | 52972 | 0.8806 | 0.6025 | 2.3014 |
| 7 | Alpha-lactalbulmin | 1A4V | 1.4 | 14.17 | 15870 | 2217 | 3926 | 6142 | 7090 | 16266 | 0.5054 | 0.4653 | 1.7686 |
| 8 | Alpha-lactalbulmin | 1A4V | 20.0 | 14.17 | 15870 | 132 | 4557 | 4690 | 20530 | 24158 | 0.8617 | 0.6004 | 1.7530 |
| 9 | Beta-lactoglobulin | 3BLG | 1.4 | 18.32 | 20518 | 2554 | 4916 | 7470 | 8179 | 21956 | 0.5076 | 0.4671 | 1.8791 |
| 10 | Beta-lactoglobulin | 3BLG | 20.0 | 18.32 | 20518 | 141 | 5276 | 5418 | 21829 | 31373 | 0.8879 | 0.6033 | 1.9133 |
| 11 | Cry1Ac protoxin | 4ARX | 1.4 | 134.90 | 151088 | 26512 | 26512 | 85337 | 85087 | 307803 | 0.2583 | 0.2058 | 1.0485 |
| 12 | Cry1Ac protoxin | 4ARX | 20.0 | 134.90 | 151088 | 435 | 27312 | 27747 | 66424 | 224437 | 0.6437 | 0.5516 | 3.4694 |
| 13 | Cytochrome c | 1HRC | 1.4 | 12.37 | 13854 | 1972 | 3507 | 5479 | 6296 | 13372 | 0.4972 | 0.4586 | 1.7338 |
| 14 | Cytochrome c | 1HRC | 20.0 | 12.37 | 13854 | 121 | 3853 | 3973 | 18590 | 20381 | 0.9082 | 0.6059 | 1.7298 |
| 15 | Fibrinogen | 1M1J | 1.4 | 311.78 | 349194 | 28068 | 59011 | 87078 | 89799 | 259562 | 0.2260 | 0.1687 | 2.2255 |
| 16 | Fibrinogen | 1M1J | 20.0 | 311.78 | 349194 | 895 | 54598 | 55493 | 136825 | 401292 | 0.4741 | 0.4386 | 4.3282 |
| 17 | Fibronectin | 3M7P | 1.4 | 550.00 | 616000 | 1021 | 1482 | 2503 | 3279 | 4693 | 0.5417 | 0.4930 | 166.5667 |
| 18 | Fibronectin | 3M7P | 20.0 | 550.00 | 616000 | 100 | 2231 | 2331 | 15283 | 7459 | 0.7919 | 0.5920 | 146.8680 |
| 19 | Glucose oxidase | 1GPE | 1.4 | 133.16 | 149139 | 11321 | 25756 | 37077 | 36256 | 147701 | 0.3645 | 0.3278 | 2.6898 |
| 20 | Glucose oxidase | 1GPE | 20.0 | 133.16 | 149139 | 319 | 19857 | 20175 | 47646 | 217450 | 0.8668 | 0.6009 | 3.8104 |
| 21 | Hemoglobin | 1BUW | 1.4 | 64.73 | 72498 | 7790 | 17594 | 25384 | 24663 | 71811 | 0.3291 | 0.2879 | 1.8575 |
| 22 | Hemoglobin | 1BUW | 20.0 | 64.73 | 72498 | 232 | 12471 | 12703 | 34847 | 115622 | 0.9035 | 0.6053 | 2.8427 |
| 23 | Human (or bovin) Serum Albumin | 1BJ5 | 1.4 | 67.71 | 75835 | 9335 | 19811 | 29146 | 29678 | 72819 | 0.2893 | 0.2418 | 1.6169 |
| 24 | Human (or bovin) Serum Albumin | 1BJ5 | 20.0 | 67.71 | 75835 | 387 | 26596 | 26983 | 61309 | 273000 | 0.7542 | 0.5856 | 1.6224 |
| 25 | Immunoglobulin G (or anti hPSA) | 1IGT | 1.4 | 148.68 | 166522 | 19877 | 39697 | 59574 | 63607 | 169383 | 0.2485 | 0.1944 | 1.6259 |
| 26 | Immunoglobulin G (or anti hPSA) | 1IGT | 20.0 | 148.68 | 166522 | 540 | 36775 | 37315 | 82973 | 322313 | 0.6092 | 0.5350 | 2.9136 |
| 27 | Immunoglobulin M | 2RCJ | 1.4 | 990.00 | 1108800 | 137060 | 175715 | 312775 | 422993 | 253122 | 0.0619 | 0.0192 | 0.8177 |
| 28 | Immunoglobulin M | 2RCJ | 20.0 | 990.00 | 1108800 | 2410 | 165764 | 168174 | 336201 | 1392551 | 0.3586 | 0.3213 | 4.3918 |
| 29 | Insulin | 1HLS | 1.4 | 5.79 | 6485 | 1257 | 1880 | 3137 | 4031 | 6107 | 0.5150 | 0.4730 | 1.4114 |
| 30 | Insulin | 1HLS | 20.0 | 5.79 | 6485 | 102 | 2546 | 2648 | 15911 | 10081 | 0.8523 | 0.5994 | 1.2805 |
| 31 | Lactoferrin | 1B0L | 1.4 | 76.52 | 85702 | 8869 | 19151 | 28020 | 28525 | 88310 | 0.3423 | 0.3029 | 2.0125 |
| 32 | Lactoferrin | 1B0L | 20.0 | 76.52 | 85702 | 271 | 16215 | 16487 | 42860 | 142619 | 0.8007 | 0.5932 | 2.8635 |
| 33 | Lysozyme | 2LYZ | 1.4 | 14.33 | 16050 | 2063 | 3734 | 5796 | 6642 | 16071 | 0.5313 | 0.4855 | 1.8810 |
| 34 | Lysozyme | 2LYZ | 20.0 | 14.33 | 16050 | 126 | 4363 | 4490 | 19965 | 23496 | 0.8836 | 0.6028 | 1.8132 |
| 35 | Myoglobin | 3RGK | 1.4 | 17.98 | 20138 | 2497 | 4719 | 7216 | 7993 | 19109 | 0.4790 | 0.4430 | 1.9188 |
| 36 | Myoglobin | 3RGK | 20.0 | 17.98 | 20138 | 138 | 5040 | 5178 | 21387 | 28331 | 0.8680 | 0.6011 | 2.0024 |
Software Specifications
- ProMS is written in Python 3
- Makes heavy use of Pandas & Tkinter modules
- Extends Schrodinger’s PyMOL as a “plugin” that can be installed via their main application
- Uses Michael Connolly’s msRoll.exe program to compute the molecular surface (part of his molecular surface package)
- Selected protein outputs have been stored in the same MySQL database as BAD

Additional Information
- For access to the ProMS source code, please contact the lab
- User manual can be downloaded here
Relevant papers & links:
- Nicolau’s paper on mapping hydrophobicity on molecular surfaces at the atomic level
- Nicolau’s paper on ProMS’ predecessor, PSPC
- Connolly’s algorithm to compute molecular surfaces analytically
- Connolly’s molecular surface package
- Schrodinger’s PyMOL